Club TAVE Team MediVision with 3D Medical SAM
3D Medical SAM with Frequency-based Dual-path Adapter
3D SAM with FDA(Frequency-based Dual-path Adapter) for Prompt-based Volumetric Medical Image Segmentation.
3DMedSAM-FDA was proposed to overcome the limitation of the original 3D SAM Adapter, which fails to sufficiently capture local structures and boundary details compared to global semantic information. We introduce a novel Dual-Path Adapter that processes global contextual information and frequency-based local features in parallel.
The detailed code can be found in the link.
Architecture
The overall segmentation framework consists of an image encoder, a prompt encoder, and a mask decoder, and follows the 3DSAM-Adapter approach, where a simple adapter is inserted into the image encoder for fine-tuning while the pretrained SAM (Segment Anything Model) is kept frozen.
Unlike the conventional single-path adapter, our framework adopts a Dual-Path Adapter composed of a Global Path and a Local Path. The detailed architecture is as follows:
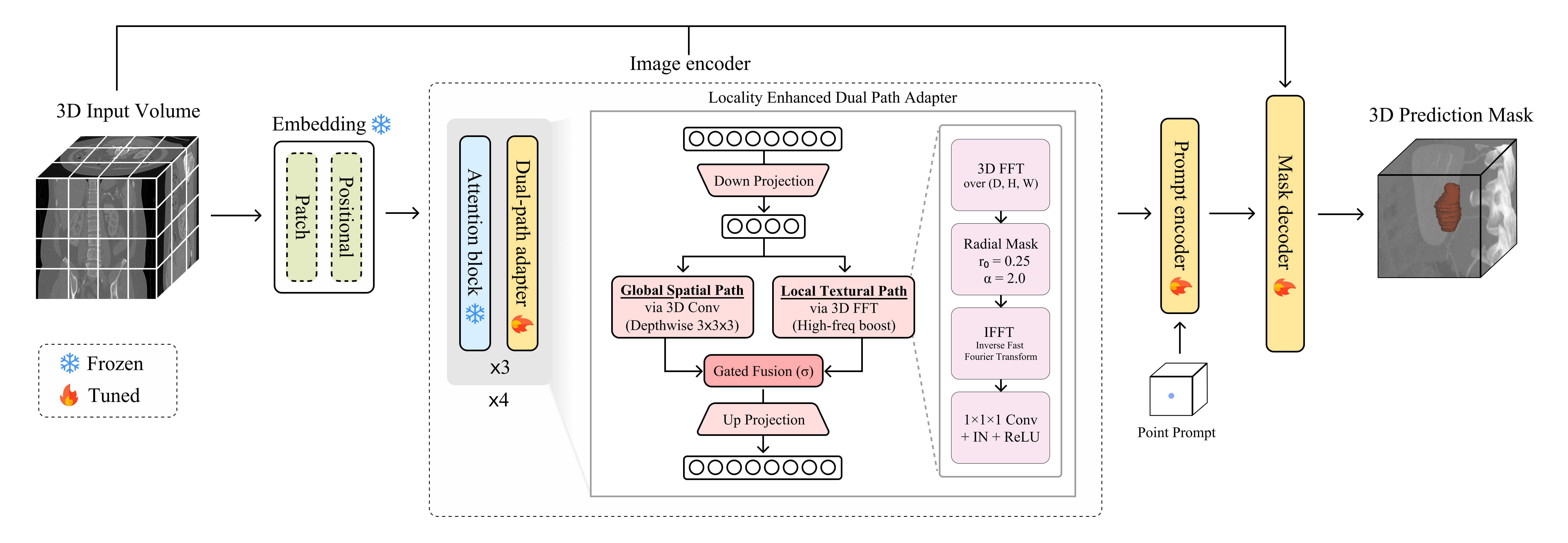
The proposed adapter (Locality-enhanced, Frequency-based Dual-path Adapter, \textbf{FDA}) operates as follows:
-
Global Context Path (Spatial Domain): A (3 \times 3 \times 3) depth-wise convolution is applied to capture 3D structural information and long-range contextual dependencies.
-
Local Textural Path (Frequency Domain):
The input features are transformed into the frequency domain using the Fast Fourier Transform (FFT).
A radial mask ( M(r) ) is applied to enhance high-frequency components (i.e., boundary information).
The features are then transformed back to the spatial domain via the Inverse FFT (IFFT), followed by a (1 \times 1 \times 1) convolution. -
Adaptive Gated Fusion:
The outputs are fused by adaptively modulating the contribution of the local path based on the global context.
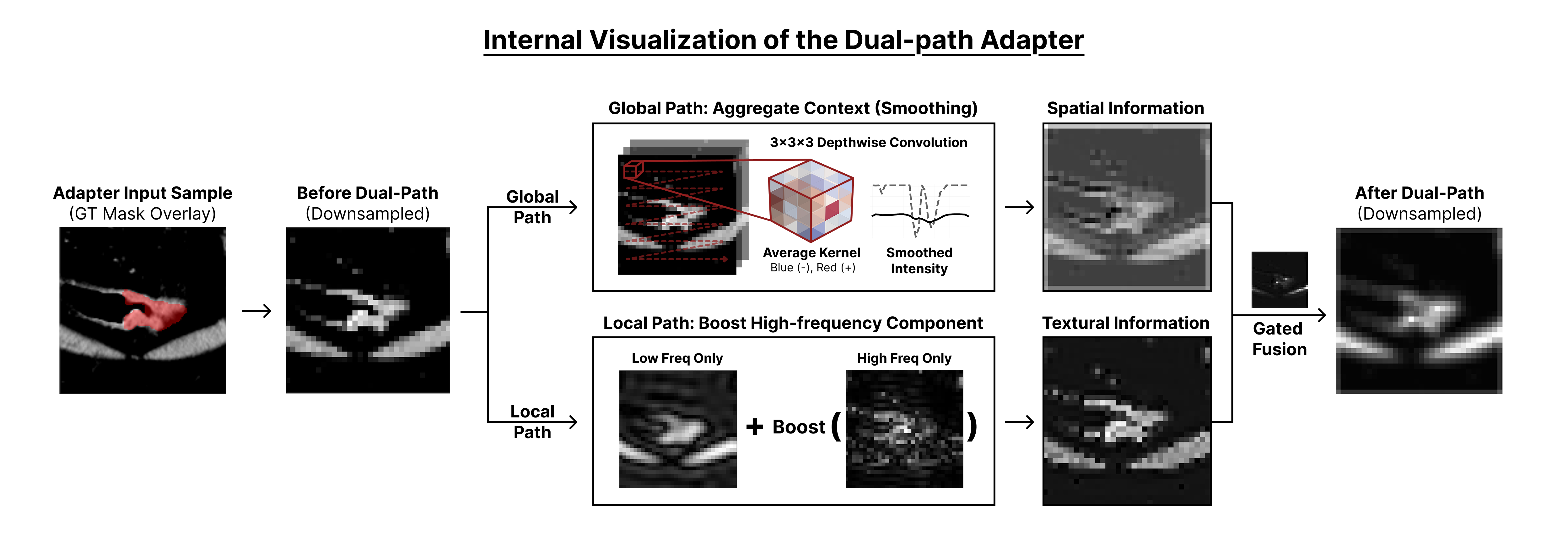
The figure above illustrates an example of the internal operation of the proposed Dual-Path Adapter.
The Global Path aggregates and smooths neighboring information using a (3 \times 3 \times 3) kernel, enabling the model to capture the overall anatomical structure and spatial context of organs.
The Local Path enhances high-frequency components in the frequency domain, emphasizing tumor boundaries and fine-grained textural details in medical images.
As a result, the complementary information from the two paths is effectively fused, producing enhanced feature maps that capture global structure while preserving sharp and detailed representations of lesions.
Results
The experiments are conducted on 3D CT images for the segmentation of four types of abdominal tumors (Kidney, Pancreas, Liver, and Colon tumors).
The detailed experimental settings, including the dataset, data preprocessing, and baseline training hyperparameters, follow those of the 3DSAM-Adapter.
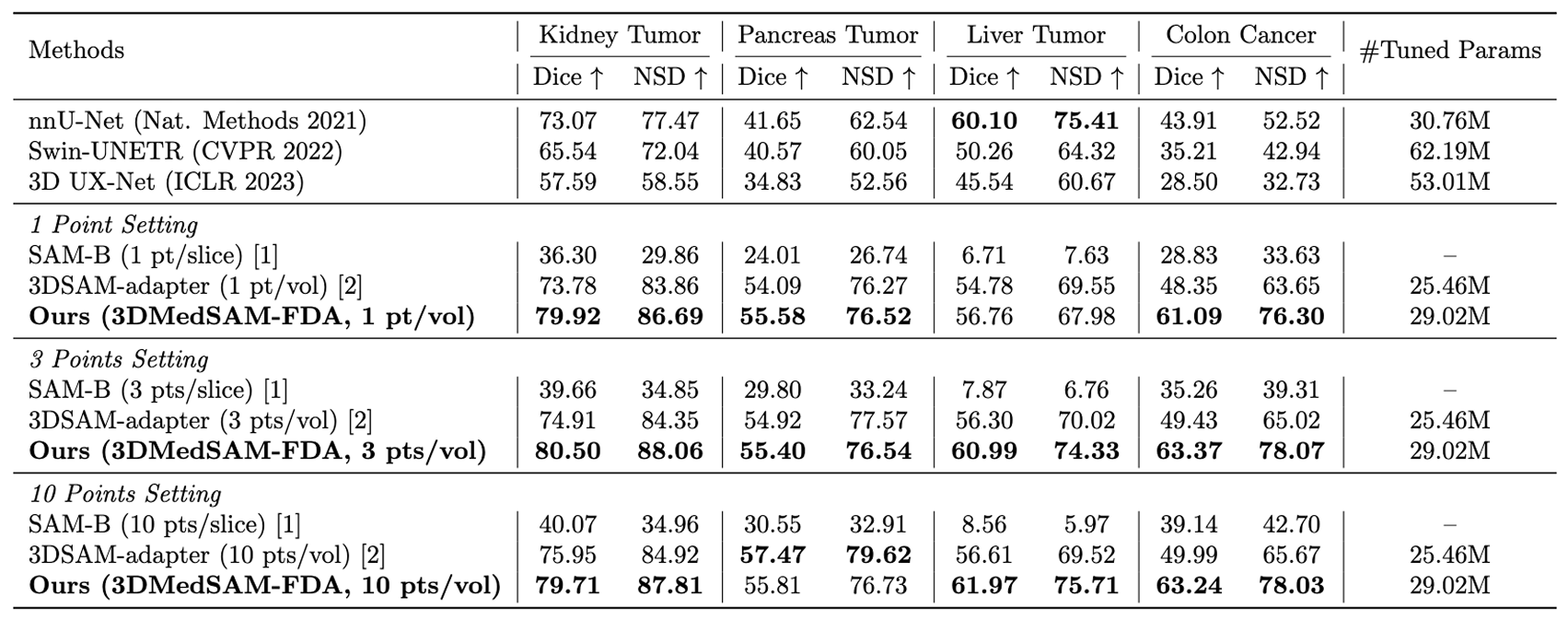
The proposed 3DMedSAM-FDA demonstrates consistent and meaningful performance improvements over the baseline across different organs. For Kidney and Pancreas tumor segmentation, it achieves average Dice score gains of +5.1%p and +0.1%p, respectively, compared to the original 3D SAM Adapter, while maintaining stable segmentation results by preserving global semantic information.
In particular, for Liver tumor segmentation, the performance improvement becomes more pronounced as the number of prompt points increases. Under the 10-point setting, our method outperforms not only the 3D SAM Adapter but also nnU-Net, achieving a +1.87%p higher Dice score. This result highlights the effectiveness of the dual-path architecture in jointly leveraging global contextual and local structural information.
For Colon tumor segmentation, our method achieves substantial performance gains over the baseline across all point settings. Even in challenging scenarios involving small and complex lesions where previous models struggle, the proposed approach consistently improves segmentation performance by enhancing boundary and fine-grained structural details through the frequency-based local path.
Moreover, the proposed model achieves these improvements while maintaining a comparable number of parameters (29.02M) to the original 3D SAM Adapter (25.46M). This demonstrates that the proposed lightweight architecture can effectively enhance representational capacity without a significant increase in model complexity.
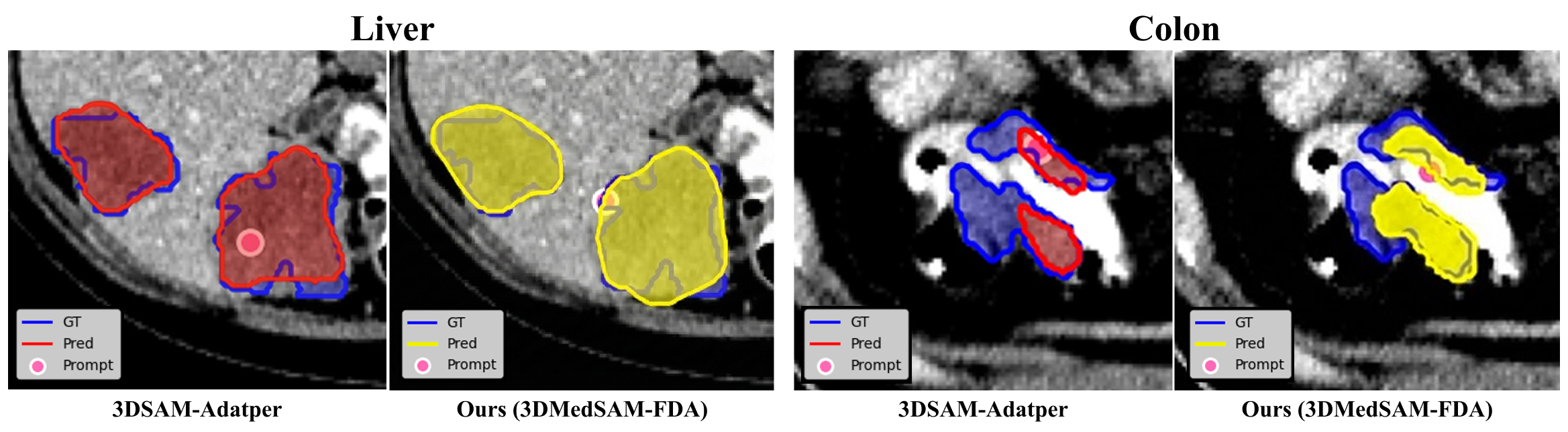
The above figure presents qualitative results, illustrating that the proposed method achieves superior performance compared to the 3D SAM Adapter, particularly for modalities where fine-grained textures and boundary details are critical, such as Liver and Colon tumor segmentation.